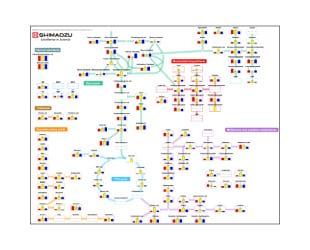
Exact Mass Database for Endogenous Metabolites
LabSolutions™ LCMS for LCMS-9030

Ready-to-use methods for metabolomics
The physical properties of target compounds in metabolomics can vary greatly. An analysis method tailored to the target compounds is therefore needed for comprehensive measurements of metabolites.
This database contains multiple ready-to-use methods for comprehensive analysis of metabolites with different properties. Pretreatment examples are also included for various metabolites and samples. The user can begin a metabolomics analysis without time-consuming evaluation of LC or MS conditions.
Features
News / Events
-
Shimadzu Scientific Instruments Inc. Opens New Laboratory in the San Francisco Bay Area -Accelerating New Product and Business Development for the Pharmaceutical and Life Science Fields
Shimadzu Corporation has positioned North America as a strategic focus area and is committed to expanding its business. Through this South San Francisco Lab, the company aims to position itself as a market leader by delivering unique, next-generation solutions that contribute to the advancement of the industry.
-
MSACL (Mass Spectrometry Applications to the Clinical Lab) 2025
September 23-25
Hotel Bonaventure Conference Center
Montreal, Canada
-
Shimadzu Announces New LCMS-8065XE Triple Quadrupole Mass Spectrometer
Shimadzu Scientific Instruments announces the LCMS-8065XE, the latest evolution in triple quadrupole mass spectrometry that redefines analytical excellence with world-class sensitivity, efficiency and sustainability.
-
Shimadzu Announces New LCMS RX Triple Quadrupole Mass Spectrometry Series
Shimadzu Scientific Instruments announces the new LCMS RX series of triple quadrupole mass spectrometry instruments. These advanced systems are supported by integrated hardware and software technologies to ensure reliable and robust results at lower operating costs, even under the most challenging and dynamic laboratory environments.
-
Innovative Technology Developed by Shimadzu and Washington University is Expected to Improve Healthy Life Expectancy
-
New Q-TOF LCMS-9050 Delivers Exceptional Mass Accuracy and Fast, Stable Polarity Switching